Autism and Genetic Testing: An Update for Clinical Practice
ABSTRACT: Routine karyotype with reflex to array comparative genomic hybridization microarray analysis (CMA) is usually recommended for children with autism spectrum disorder (ASD) who have no findings of recognizable syndromes that can be diagnosed by individual fluorescence in situ hybridization tests. It is important to explain to parents considering genetic testing that a positive result indicates an under-lying condition associated with autistic behaviors not the presence, severity, or type of ASD. Pediatricians can help families decide about subspecialty referral and testing through a frank discussion of their needs based on physical examination findings and neurodevelopmental assessment. CMA may be useful in children with ASD and significant developmental delay; however, parents must be informed that the results will not confirm or classify their child’s autism.
Beyond the crucial role of screening for autistic behaviors is the opportunity for the pediatrician to help explain genetic findings or diagnoses and integrate this information into the medical home. Research shows that autism has multiple genetic and nongenetic causes and that no particular genetic or nongenetic cause is linked to a specific autism spectrum disorder (ASD).1 Twin studies indicate a variable genetic contribution to autism, shown by a concordance of 58% to 100% for monozygotic twins (higher for female twins), 31% for dizygotic twins, and 5% to 22% for siblings.2,3 Common environmental factors among affected children include prematurity (27%) and hearing or vision loss (7% to 25%)2,3; congenital influences—rubella infection or thalidomide exposure—account for a few cases.1
For the child with associated symptoms (eg, seizures and dysmorphology), pediatricians can consider referral to the neurologist or geneticist. They can assist in the formal diagnosis of autism by reviewing disability assessments (eg, Autism Diagnostic Observation Schedule and Autism Diagnostic Interview-Revised) with the developmental pediatric specialist or psychiatrist. They must also remain cognizant of underlying diagnoses— determined through birth history, global developmental delay, or developmental regression (such as Williams syndrome and Rett syndrome)—that dictate additional preventive health care.4
A recent dilemma for primary physicians and their patients with autism is the new technology of array-comparative genomic hybridization (aCGH, microarray analysis, CGH-microarray analysis or CMA). The yield of positive results in CMA is as high as 22% in some studies, compared with 2% to 3% for routine karyotyping; however, normal variations (copy number variants or CNVs) in the parents reduce the yield of significant CMA findings to 7% to 10%.5-7 Because of the greater emphasis on genetic testing, including CMA, in autism, we review its use through 3 clinical case presentations.
AUTISM DEFINITION
AND DIAGNOSIS
Autism was first described by Kanner in 1943 as 3 types of behaviors: abnormal verbal and nonverbal communication, abnormal social interaction, and repetitive movements and routines.8 Asperger in 1944 described children with motor and social immaturity, poor nonverbal communication, eccentric behaviors with restricted areas of interest, yet normal language development and intelligence.9,10 Asperger syndrome, pervasive developmental disorder not otherwise specified, and autism disorders are now often grouped as ASD.9,10 Autistic behaviors can occur in most disorders with cognitive disability, ranging from genetic disorders (fragile X or Down syndrome) to conditions of the maternofetal environment, such as fetal alcohol spectrum disorder. The term “autism” should be viewed as a heterogenous spectrum that, like diabetes or schizophrenia, follows the inheritance model of multifactorial determination.3
ASD is classically diagnosed at ages 3 to 4 years, when deficits in communication and socialization and repetitive movements and restricted interests are brought to the physician’s attention.4 Earlier recognition of autism would allow timely employment of effective therapies, such as applied behavioral analysis.6 Diagnosis by 18 months through increased use of screening questionnaires such as the Modified Checklist for Autism in Toddlers, which evaluates eye contact, pointing, bringing things to parents, hypersensitivities to loud noises or food textures (see www.m-chat.org), is now emphasized by the American Academy of Pediatrics.4 Early diagnosis could take advantage of neuroplasticity before critical periods expire; it has been shown that visual pathways can be remodeled until age 2 years by stimulating the deviant eye.11 However, no reliable or valid diagnostic tool for ASD is available until the second year of life. Early clinical recognition of ASD remains in the hands of pediatric and family practitioners along with ancillary health professionals.
TYPES OF GENETIC TESTS
Routine karyotype ( about $450) with reflex to CMA ($1800)—reflex, meaning proceed to the next test if the prior one is normal—is usually recommended for children with ASD who have no findings of recognizable syndromes that can be diagnosed by individual fluorescence
in situ hybridization (FISH) tests (about $200). Specific DNA tests for fragile X syndrome (in boys) or Rett syndrome (in girls) may be added, because CMA cannot detect changes at the gene level (eg, expanded triplet repeats or single nucleotide mutations). Most DNA tests cost $2000 to $3000.
 PRETEST COUNSELING
PRETEST COUNSELING
AND PLANNING
It is important to explain to parents that a positive genetic test result indicates an underlying condition associated with autistic behaviors not the presence, severity, or type of ASD. The clinician’s dysmorphology skills often dictate whether the more expensive CMA test is used rather than a specific FISH test. Many laboratories advocate first-line use of CMA without prior karyotyping because it will screen for all of the standard FISH deletions or duplications (eg, deletion/duplication of chromosome 5 in 10% of patients with Sotos syndrome, duplication of chromosome 17 in a form of Charcot-Marie-Tooth disease).12
For children with ASD who are not dysmorphic or have no specific neurological findings, which is often the case, it is prudent to enlist the help of a geneticist and/or neurologist to guide testing. The neurologist is helpful when the child requires additional neuromuscular tests or imaging, the geneticist can provide guidance when unusual CMA variants or normal results dictate consideration of further tests (eg, DNA tests for Noonan or other syndromes). CMA will not detect balanced chromosome translocations or low levels of mosaicism, boys with high-functioning autism will be unlikely to have fragile X syndrome, and girls without regression, hand-wringing movements, irregular breathing, or circulatory changes are unlikely to have Rett syndrome.
Although routine karyotype with reflex to CMA (and fragile X in boys) can be justified13 and is usually covered by insurance, good medicine dictates careful coordination of clinical and laboratory evaluations by involving subspecialty clinicians. Pretest planning is particularly important as presymptomatic diagnosis (eg, Huntington disease, breast-ovarian cancer susceptibility) becomes more common and children with such a diagnosis face potential scholastic, insurance, or employment discrimination, despite the enactment of the Genetic Information Nondiscrimination Act.14
WHEN GENETIC TESTING INDICATES
AUTISM SUSCEPTIBILITY
Routine karyotype. The routine karyotype is generally ordered for children with growth/developmental delays and dysmorphology exemplified by the infant in Case 1. This test is performed on lymphocytes from anticoagulated blood (sodium heparin green-top tube stored at room temperature). The white cells are separated, cultured, arrested in metaphase, analyzed by inspection, and arranged by computer to form a karyotype (Figure 1). Results are available within 10 to 14 days.
In Case 1, the diagnosis of Wolf-Hirschhorn syndrome guided preventive health care, with surveillance for eye, cardiac, and renal anomalies as well as for ASD. The risk of autism is increased in Wolf-Hirschhorn syndrome, as in all chromosome disorders, and this risk increases with the degree of mental disability (the risk of ASD in Down syndrome is 10%). Early surveillance for ASD initially focuses on learning differences associated with chromosome disorders and later may lead to more directed assessments for autistic behaviors.
An important benefit of genetic diagnosis is that it helps coordinate genetic counseling and testing of the parents. Results of parental studies (for chromosome rearrangements, such as translocation, deletion, and duplication) in Case 1 were normal, with minimal risk of recurrence.
FISH testing. As subtle chromosome changes were recognized by improved banding techniques, fluorescently labeled DNA segments (FISH probes) were developed to target potential deletion or duplication regions. Deletion of a chromosome band could be more easily recognized by the lack of fluorescent signal, compared to its partner chromosome (homologue). Thus, the routine karyotype was supplemented with a specific FISH test that targeted the region of interest (eg, chromosome 7 in Williams syndrome, see Case 2). Multiple FISH probes were then used to screen patients for several conditions (eg, 7q deletions in Williams, 17p in Smith-Magenis, 22q in DiGeorge/velocardiofacial syndromes).
More than 40 DNA probes are used with FISH to detect common changes near the chromosome ends (subtelomere FISH). This technique will be replaced by CMA, which has 44,000 to 1 million probes on a DNA chip. Use of the rapid FISH panel, which has probes directed at chromosomes 13, 18, 21, X, and Y, is likely to continue. This test allows 24-hour diagnoses of trisomy 13, trisomy 18, Down, and Turner syndromes for timely prenatal/neonatal management.
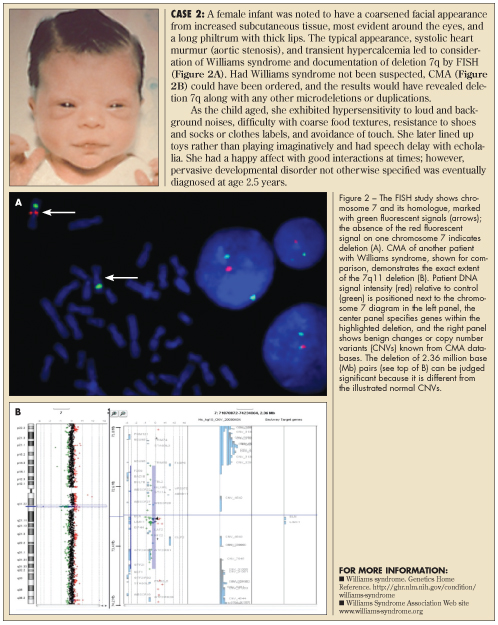
The infant in Case 2 had a specific FISH test for 7p11 deletion (Figure 2A) because genetic and cardiology consultants suggested the diagnosis of Williams syndrome. Most common microdeletions, such as those of Williams, Prader-Willi, and velocardiofacial syndromes, are adequately defined by FISH. When the diagnosis of Williams syndrome is unclear or the findings do not point to a single recognizable syndrome, CMA is often considered because it can detect deletions anywhere in the genome without directing the laboratory’s attention to any particular chromosome region. CMA is also useful as a research tool in that it can define precisely which genes are duplicated or deleted and improve correlations between clinical syndromes and cytogenetic changes.
The pediatrician’s role in Case 2 is to reinforce subspecialty counseling about Williams syndrome and review its variable intellectual outcome (IQ of 40 to 80), happy personality, and minimal recurrence risks for the family. Parental studies could be considered in rare situations in which parents have symptoms of Williams syndrome. Risks of autism and other medical complications may be mentioned initially then oriented towards specific symptoms as development and early intervention progressed. Because as many as 50% of young children with Williams syndrome show signs of ASD, surveillance for autistic behaviors should be combined with that for strabismus, heart defects, hypertension, and renal disease in this infant.15
 GENETIC TESTING IN CHILDREN WITH
GENETIC TESTING IN CHILDREN WITH
AN ASD DIAGNOSIS
One benefit of genetic testing in children with ASD is to identify a possible genetic cause, which can alleviate inappropriate maternal guilt and define recurrence risks for future pregnancies. Another more important benefit in young children is to recognize medical complications associated with the genetic condition and institute preventive health care. Claims that a positive genetic test result can influence therapy or management for children with minimal dysmorphology or birth defects are less convincing;12 even prognosis is often difficult, as discussed in Case 3.
Pediatricians can help families decide about subspecialty referral and testing through a frank discussion of their needs based on physical examination findings and neurodevelopmental assessment. CMA may be useful in children with ASD and significant developmental delay; however, parents must be informed that the results will not confirm or classify their child’s autism.
CMA technology. The CMA technique compares patient and control hybridization to hundreds of thousands of DNA segments on a grid (array) that shows subtle extra (duplicated) or missing (deleted) chromosome material anywhere on the 23 + X or Y chromosomes. CMA can be visualized as the equivalent of more than a million simultaneous FISH tests, but in CMA, the DNA segment probes are arranged on a grid, and patient and control DNA are tagged with fluorescence.5-7 Patient DNA is isolated from EDTA-anticoagulated blood (lavender-top tube) and tagged with one fluorescent color, while a control sample is tagged with a different color (see Figures 2B and 3). A machine graphs the signal intensities of control versus patient hybridization over all DNA segments, preserving their order along chromosomes 1-22, X, and Y. Use of CMA is increasing because it can find subtle chromosome changes in children who lack the typical dysmorphology, growth delay, or birth defects of classical chromosome disorders that are apparent in Cases 1 and 2.
For the boy in Case 3, results of routine karyotype and fragile X DNA analysis were normal; however, CMA demonstrated a deletion of significant size at band 16p11.2 (Figure 3). Reports of this same deletion in other patients with ASD support a causal relationship for this chromosome change.16
The parents could be counseled that the 16p11.2 deletion in their child had caused an underlying disorder with increased susceptibility for ASD. Results of their FISH studies were normal, indicating that they (and their daughter) were not at increased risk for this deletion in future pregnancies.
A frequent problem with CMA is that many microdeletions/microduplications are unique or rarely encountered. Criteria are emerging to decide whether an unusual variant is significant. These include negative parental studies, sufficient size (greater than 0.5 Mb), substantive gene content of the altered chromosome material, and absence from repositories of benign variants.
Many laboratories report rare findings as “variants of unknown clinical significance.” This is temporary strategy that will hopefully be revised as CMA databases enlarge. Clinical variation among affected persons is the rule with subtle CMA findings,17and public databases like DECIPHER (http://decipher.sanger.ac.uk), UCSC Genome Bioinformatics (http://genome.ucsc.edu), and HGNC (www.genenames.org) may eventually define the clinical spectrum of underlying disorders and quantify their risks for ASD. The pediatrician can explain the reasons for prognostic uncertainty and collaborate with their preferred developmental specialist in providing ongoing assessment with sculpting of therapy and school programs. ■
References
1. Muhle R, Trentacoste SV, Rapin I. The genetics of autism. Pediatrics. 2004;113:e472-e486.
2. Hallmayer J, Cleveland S, Torres A, et al. Genetic heritability and shared environmental factors among twin pairs with autism. Arch Gen Psychiatry. 2011 Jul 4. [Epub ahead of print].
3. Rosenberg RE, Law JK, Yenokyan G, et al. Characteristics and concordance of autism spectrum disorders among 277 twin pairs. Arch Pediatr Adolesc Med. 2009 163:907-914.
4. Greenspan SI, Brazelton TB, Cordero J, et al. Guidelines for early identification, screening, and clinical management of children with autism spectrum disorders. Pediatrics. 2008;121:828-830.
5. Sagoo GS, Butterworth AS, Sanderson S, et al. Array CGH in patients with learning disability (mental retardation) and congenital anomalies: updated systematic review and meta-analysis of 19 studies and 13.926 subjects. Genet Med. 2009;11:139-146.
6. Coulter ME, Miller DT, Harris DJ, et al. Chromosomal microarray testing influences medical management. Genet Med. 2011;13:770-776.
7. Tonk VS, Wilson GN. Autism spectrum disorder with 10q26 microdeletion by subtelomere FISH. Pediatric Health, Medicine and Therapeutics. 2011;2:49-53. www.dovepress.com/autism-spectrum-disorder-with-microdeletion-10q26-by-subtelomere-fish-peer-reviewed-article-PHMT.
8. Kanner L. Autistic disturbances of affective contact. Nervous Child. 1943;2:217.
9. American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders, Fourth Edition. Arlington VA: American Psychiatric Publishing; 2000.
10. Howlin P, Magiati I, Charman T. Systematic review of early intensive behavioral interventions for children with autism. Am J Intellect Dev Disabil. 2009;114:23-41.
11. Altemeier WA, Altemeier L. How can early intensive training help a genetic disorder? Pediatr Ann. 2009;38:167-170, 172.
12. Miller DT, Adam MP, Aradhya S, et al. Consensus statement: chromosomal microarray is a first-tier clinical diagnostic test for individuals with developmental disabilities or congenital anomalies. Am J Hum Genet. 2010;86:749-764.
13. Centers for Disease Control and Prevention. Autism Spectrum Disorders (ASDs). Recommendations and Guidelines. www.cdc.gov/ncbddd/autism/hcp-recommendations.html. Accessed September 16, 2011.
14. Genetic Information Nondiscrimination Act Legislation. Genetic Alliance Web site. www.geneticalliance.org/legislation.gina. Accessed September 12, 2011.
15. Klein-Tasman BP, Phillips KD, Lord C, et al. Overlap with the autism spectrum in young children with Williams syndrome. J Dev Behav Pediatr. 2009;30:289-299.
16. Weiss LA, Shen Y, Korn JM, et al. Association between microdeletion and microduplication at 16p11.2 and autism. N Engl J Med. 2008;358:667-675.
17. Ben-Schachar S, Lanpher B, German JR, et al. Microdeletion 15q13.3: a locus with incomplete penetrance for autism, mental retardation, and psychiatric disorders. J Med Genet.2009;46:382-388.


